Automating Bumblebee Identification with Machine Learning

Written by John Farrell, Senior Developer
Identifying a Problem
As Instrument employees, we receive a budget each year for Professional Development. It’s an open-ended program, intending to support “skill building, inspiration, and the expansion of knowledge” within the studio. Michael Bunsen and I, both Senior Developers, were taking some machine learning classes outside of work, so it felt like a natural topic to explore for a Professional Development project.
Machine Learning is a broad topic, so we focused our areas of interest around ecology, conservation, and cartography. A serendipitous bike ride introduced us to the Xerces Society. The Xerces Society is a nonprofit organization headquartered in Portland, OR, focused on the conservation of various invertebrate species. We connected with Rich Hatfield, Senior Conservation Biologist. “I’ve been hoping someone with machine learning experience would approach,” he told us.
Rich heads up Bumble Bee Watch, an online citizen-science platform focused on bumble bee conservation. Users record bumble bee sightings with photos, location, and attempted species identification. For every submission, an expert like Rich manually examines and verifies the species identification to ensure data accuracy. Michael and I thought this is exactly the type of problem machine learning is suited for. An automated system for identifying bee species would save Rich and his team hundreds of hours of time.
Exploring Bee Data
The team at Xerces were eager to explore a partnership and sent over their dataset of bee sightings. This included 53,308 independent sightings, of which 32,388 were already verified by experts. Those sightings often contained multiple images, which gave us a total of 72,192 labeled photographs to work with, a fantastic foundation for training an image classification system.
We started exploring our data in code by using Python and Jupyter notebooks. We received two datasets from Rich -- one with labeled photographs and another with metadata about each sighting. Each row contained an observation ID, which we used to join the two datasets together and associate each photograph with its species label.

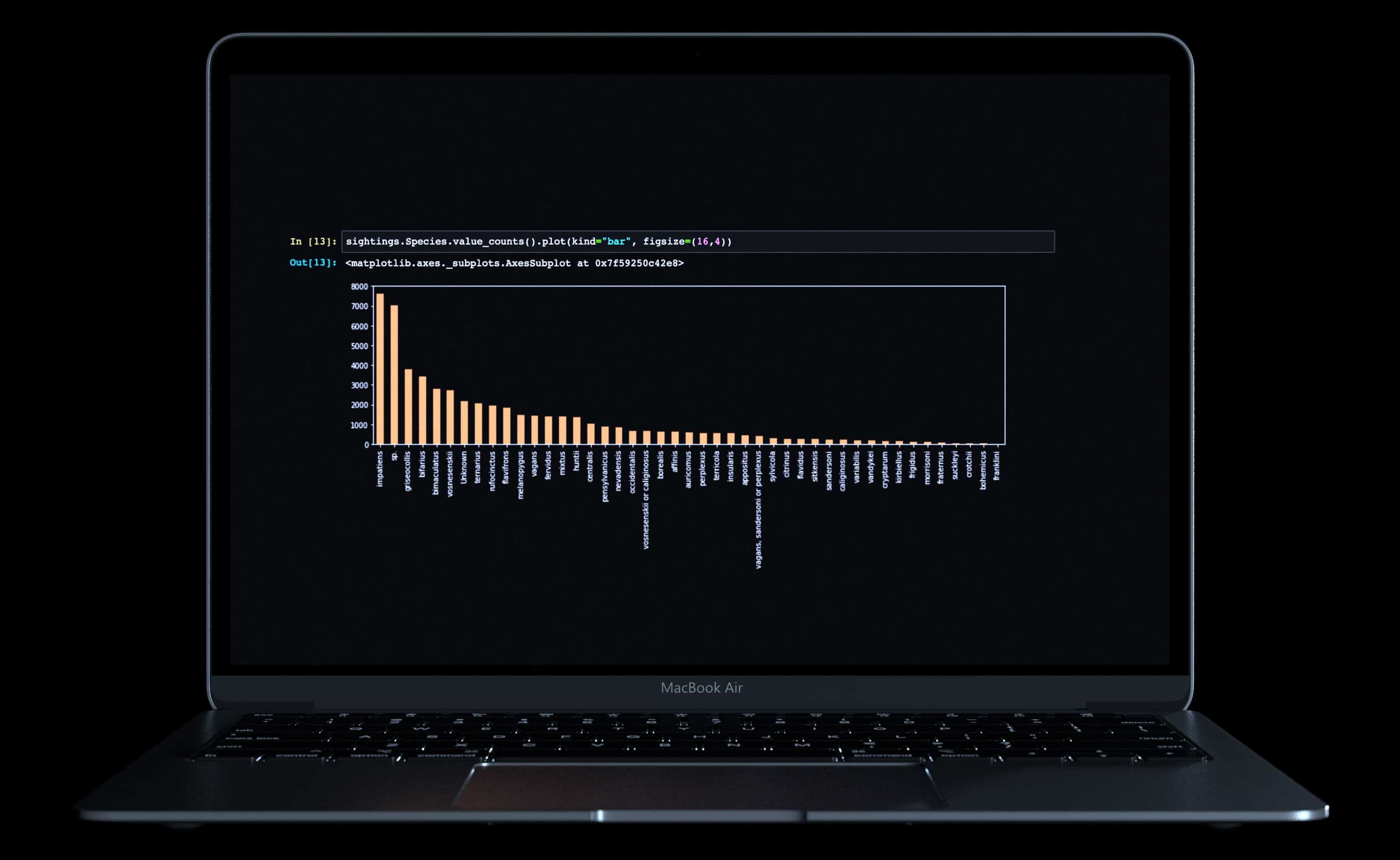
The labeled photos below consisted of the species name and photograph. There were 45 unique species in the dataset, following a long-tail distribution (including a couple outliers).
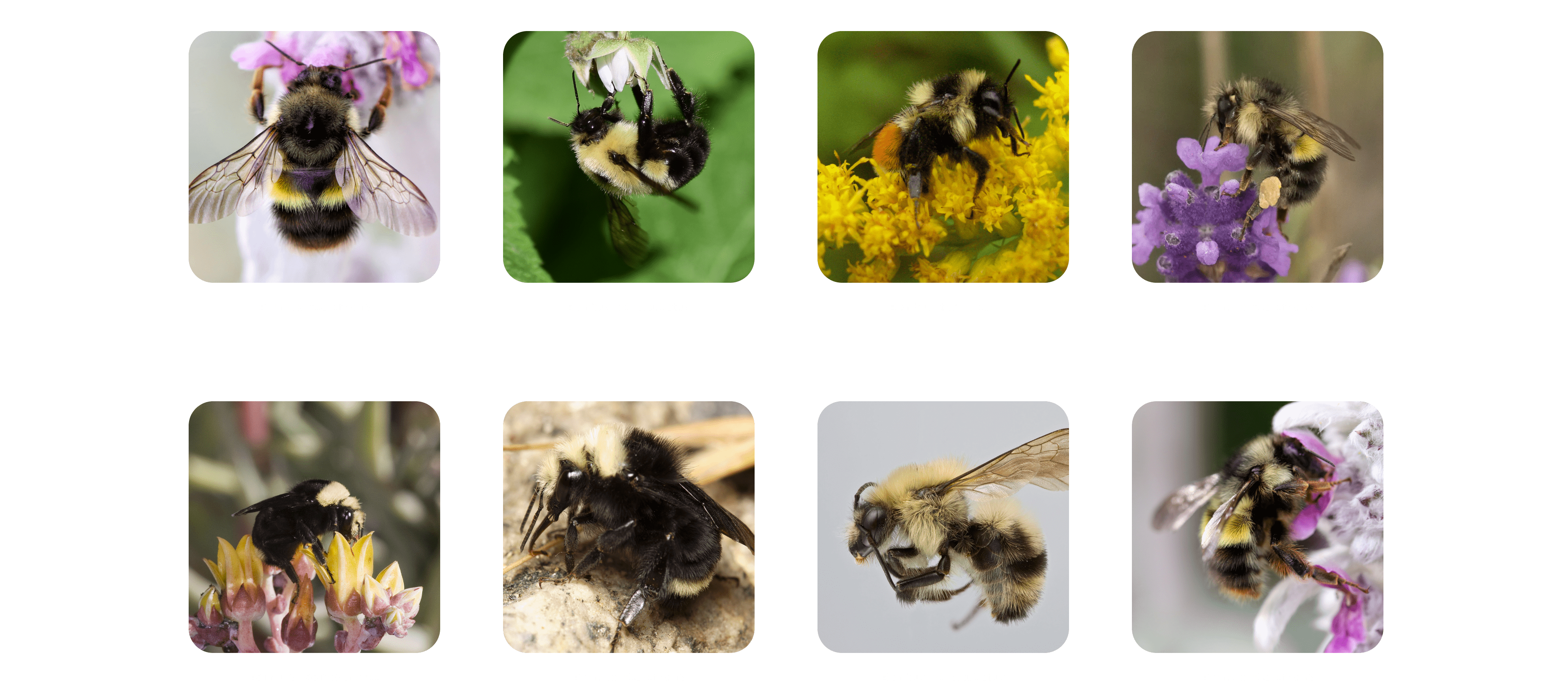

To achieve meaningfully accurate training results, we needed a minimum threshold of observations. After some experimentation, we found this threshold to be about 300 observations. Below that, the prediction accuracy was not high enough to be useful. We removed all species with an observation total under 300, and arrived at a final list of 27 species.

Next we divided the data into sets necessary for training and testing results. We used an 80/20 split for the training and validation sets, reserving 0.5% for a test set (so actually 19.5% for the validation set). This is a common ratio used to split up your data. You need the majority of the data for model training and experimentation. Some of the data is saved to validate your experimentations, and a very small amount is your reserve.
The largest set, training, is the data we used to train the model. This is the data the model “learns” with and is used to calculate the loss and gradients during the training loop. The validation set is used to test the model after each epoch of training and gives feedback about how the model is performing. The validation output impacts our decisions during training, but the model does not learn from it or directly. Test is a fresh set of data reserved to periodically test the model. It should be a representative sample of data that the model has never seen before, mimicking real-world usage.
Our final sets worked out to 57,382 training photos, 14,424 validation, and 357 for test. Storing our images in folders labeled with the species name made it very straightforward to load the images into the model.
/training
/impatiens
/bifarius
/griseocollis
image0001.jpg
image0002.jpg
image0003.jpgTraining Our Model
We started training using the FastAI framework, which is built on top of PyTorch. FastAI allowed us to work quickly as we began the project, while maintaining the flexibility and power of PyTorch to customize our training as necessary. We are utilizing residual neural networks (ResNets) architecture for training, our model uses the resnet34 architecture (34 layers to the network), with weights pre-trained on the imagenet dataset.
Each layer of the model can identify progressively more specific features. For example, the first layer may be identifying a sloped line. The next layer identifies a curve, and so on. Since these foundational learnings are consistent across image recognition applications, we can begin by using a model that has already been trained in these broad areas. We then train the final few layers on our specific application - in this case, bumblebees. This is an example of loading our dataset into the model, and running it through a training cycle.
from fastai.vision import *
from fastai.metrics import error_rate
from fastai.version import __version__
__version__
# '1.0.52'
data = ImageDataBunch.from_folder(
"data/sorted/",
size=224,
bs=16 # Batch size
# ds_tfms=get_transforms() # add random rotations, etc?
).normalize(imagenet_stats)
data.show_batch(rows=3, figsize=(7,6))
learn.fit_one_cycle(
8,
slice(7.59E-07, le-04),
callbacks=[SaveModelCallback(learn, every="improvement", monitor="error_rate")]
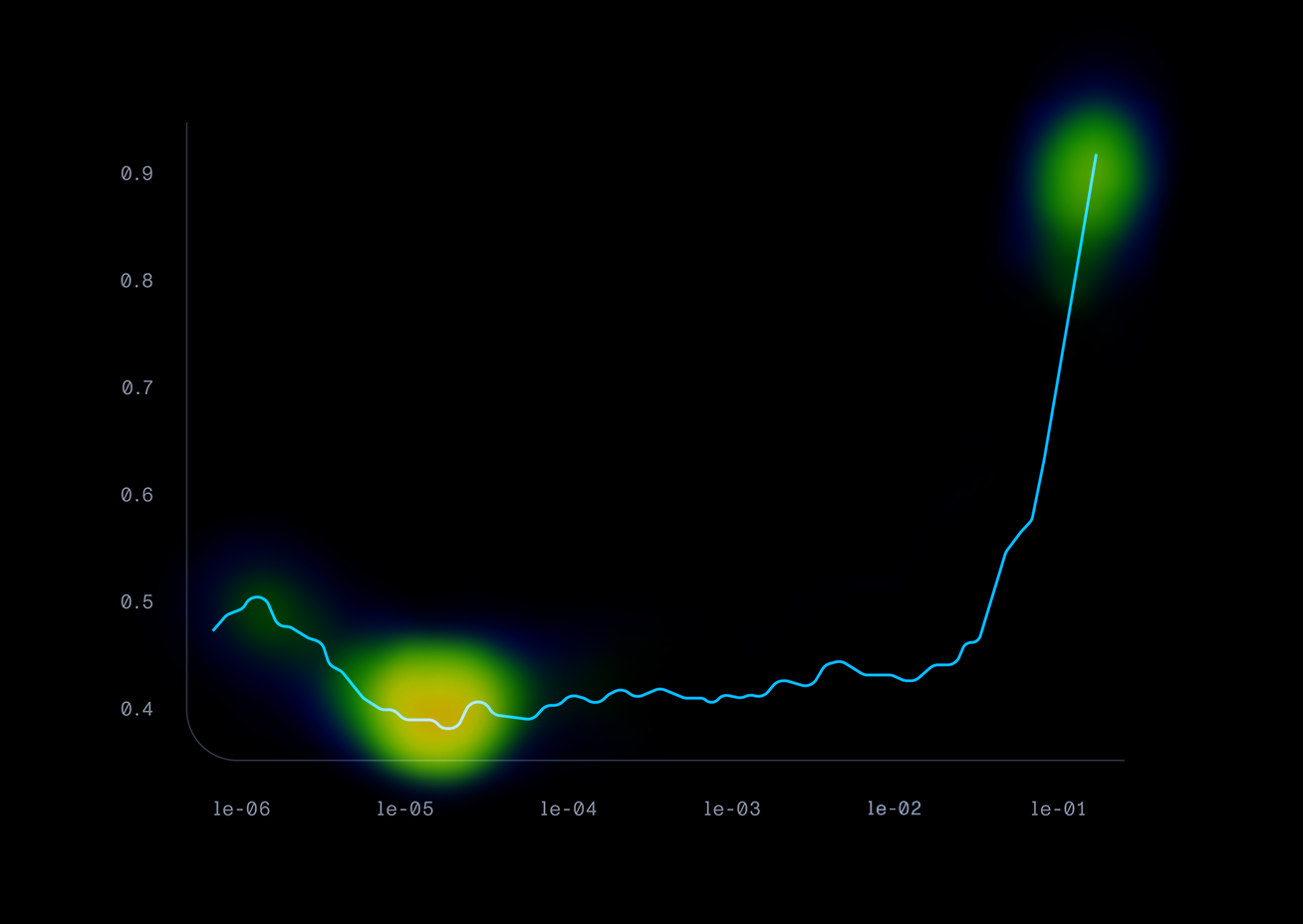
)This style of development is much different than we are used to in our day jobs. It involves many cycles of experimentation and tuning hyperparameters, rather than continually writing a process flow of code. One of our methods of experimentation was “spoon-feeding” the model, where we continually trained the model on small batches of high-quality data at optimal learning rates. This was a really successful tactic to increase our model accuracy. Plotting the learning rates over time and finding the steepest descent of the curve was really useful in this process. This approach showed promising results in a decreasing error rate as we trained the model.
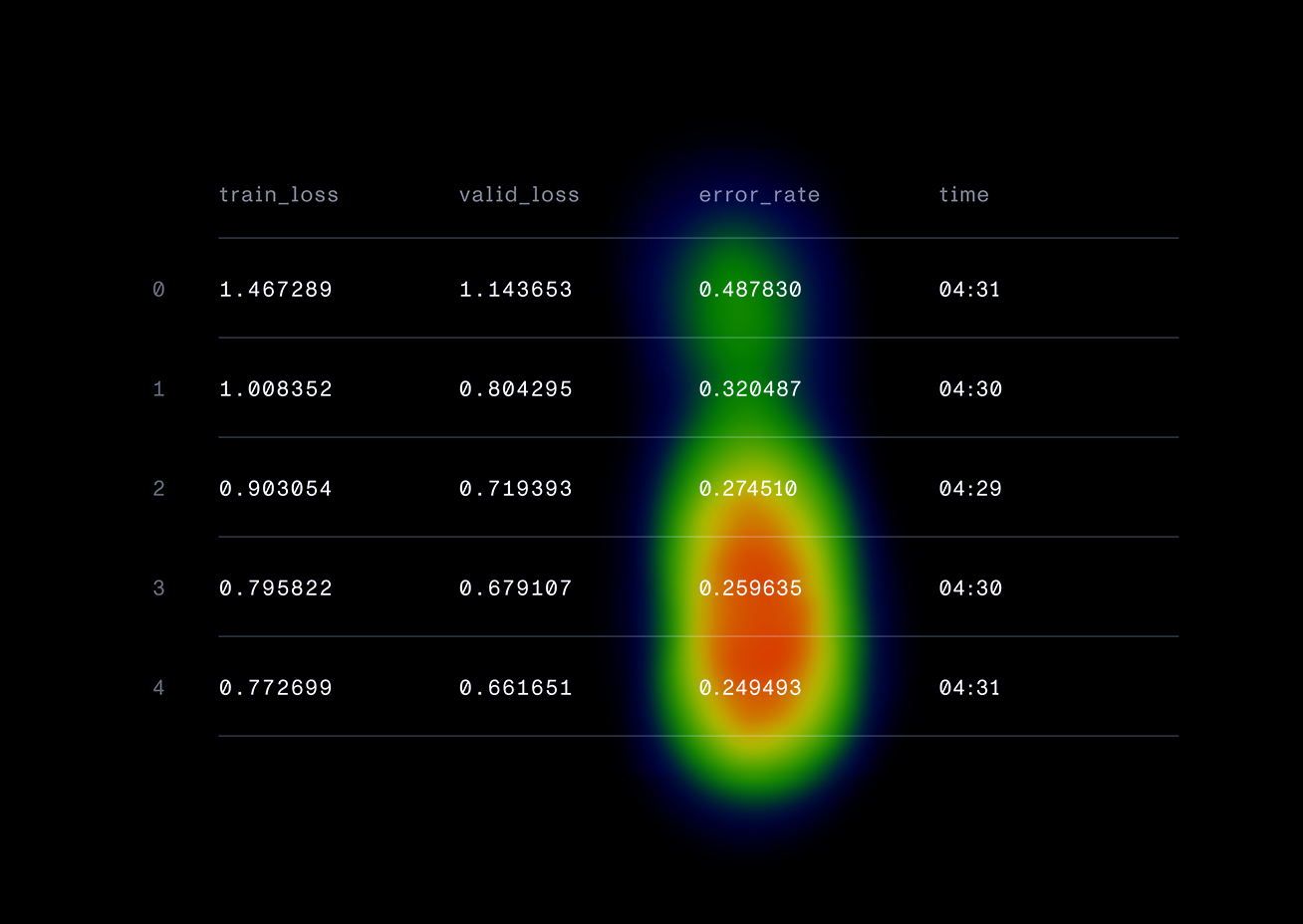
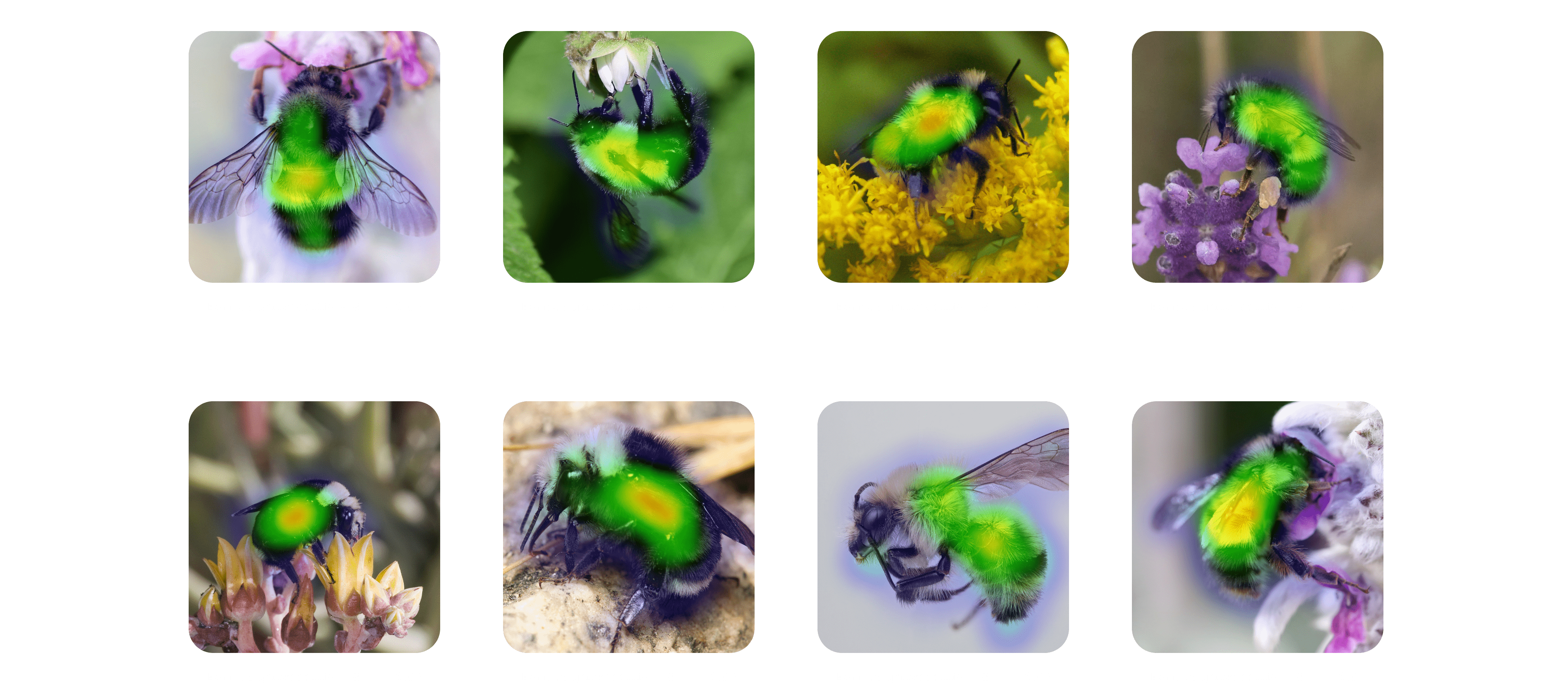
One feature of FastAI we really enjoyed was the plot_top_losses function. This means we could view the exact pixels that activated our model, meaning the areas that had the highest influence on the prediction. This plot demonstrates nine successfully classified bees - the predicted species, actual species, loss, and probability of the prediction.

Ultimately, our model achieved a success rate of 85% on species predictions. The machine learning model was only one part of our project plan. The other part of our plan after getting 85% accuracy, was to be able to push it further by more seasoned scholars, grant funding, etc. Our project plan involved a complete integration - the ability to affordably and effectively host a model and use it in a production web application to gain efficiency for expert researchers.
System Integration
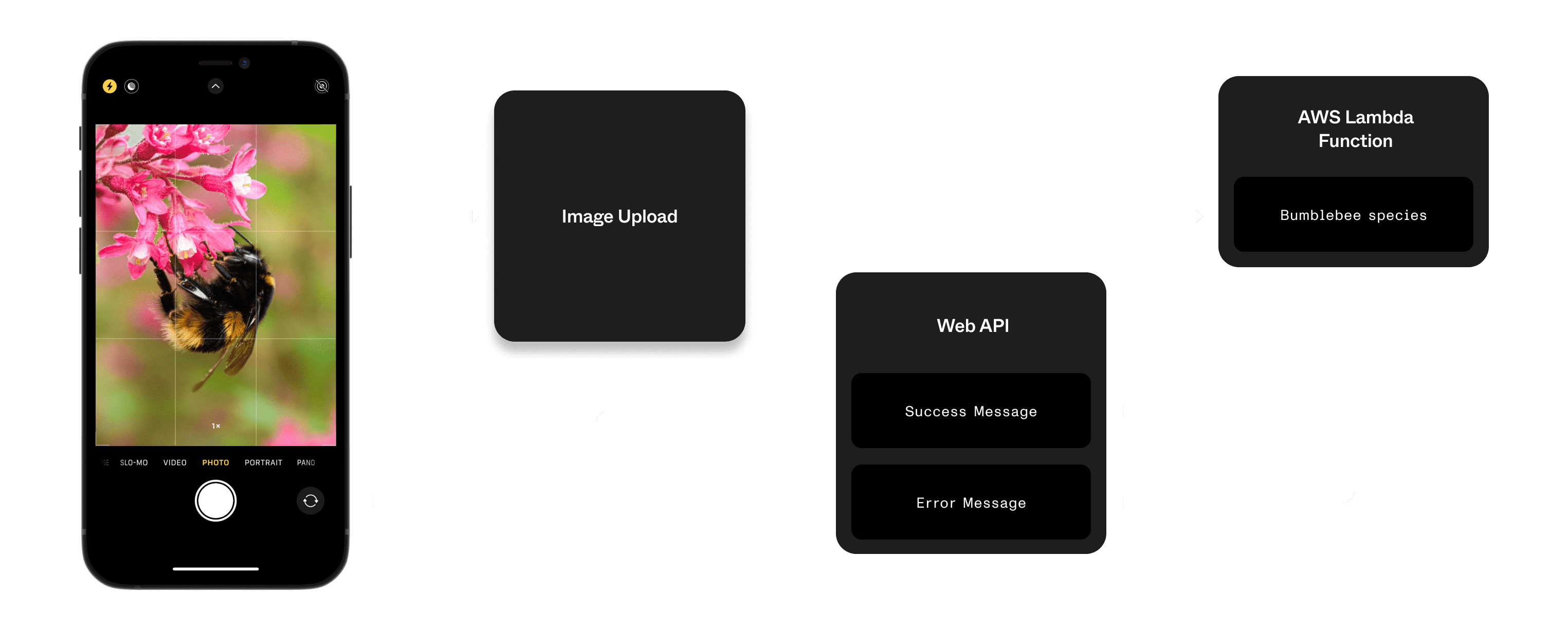
Our architecture involved hosting the model behind an AWS Lambda function within a Flask server, which we could interface with via AJAX requests. The request accepts a photograph, and returns a JSON object with a species prediction, and percentage of confidence.
{
“confidence”: 0.98712475,
“label”: “grisceocollis”
}An ideal future state of the web application passed a photograph through several models, saved results with observation records in the database, and returned various messages back to the user when they submitted an observation.
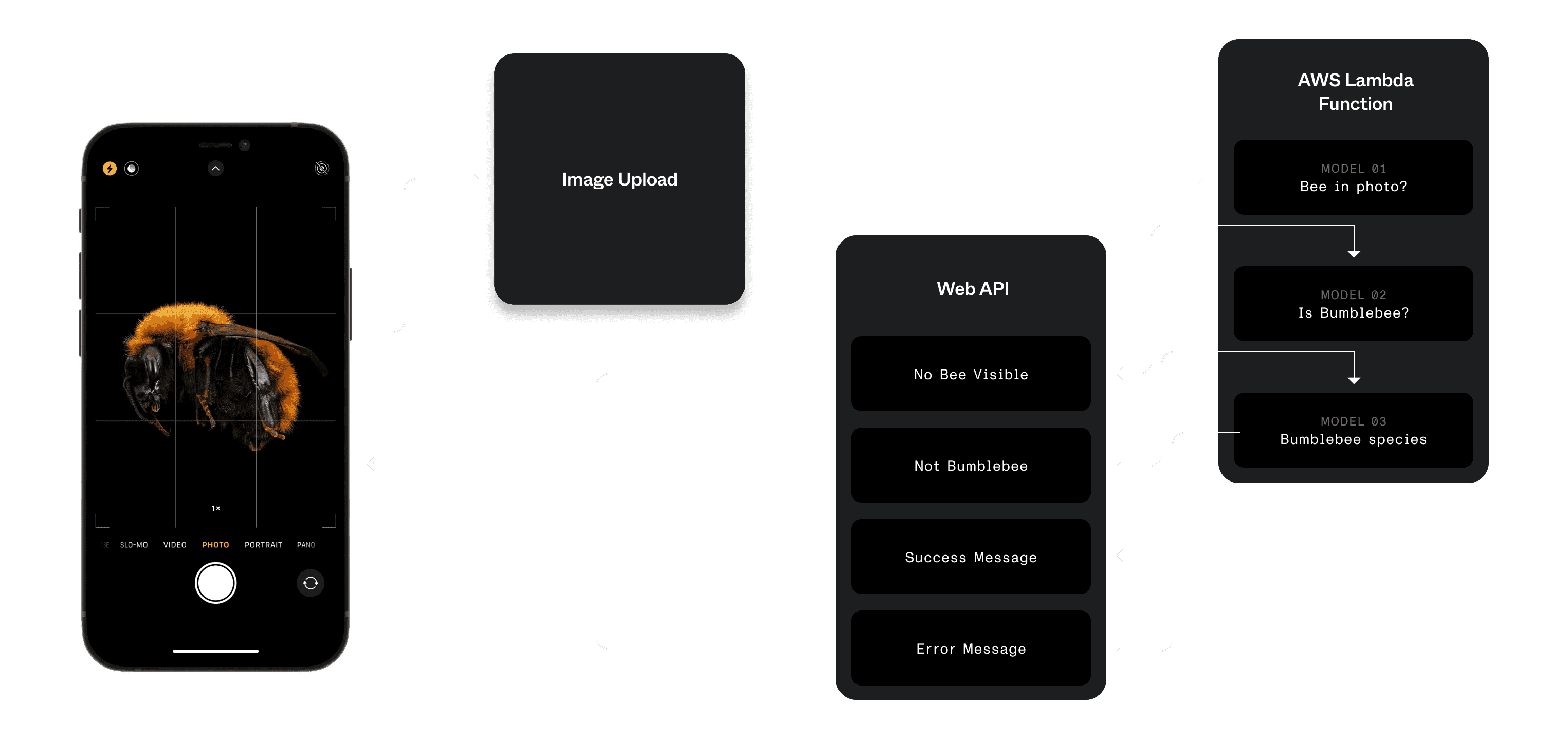
With this as the future vision, we worked towards a lighter approach so the team could get acclimated to the model. This approach would predict the bee species, and display the results only to site administrators.
When an administrator verifies a submission, they view the high-res photos. Since administrators needed to make their own expert judgements without bias, we hid the results of the model prediction behind a single button click. The button pulls the images, sends them to the classifier, and displays results to compare with expert findings.

In addition to setting up our hosted model, we wrote a vanilla Javascript class to interface with the model. We handed this code off to the developers of Bumble Bee Watch for integration. Admins of the site are now using the verification tool in their daily work, with positive early results.
/**
* @desc Send photo to Image Classifier hosted on AWS Lambda
* This should happen when a successful response comes back from image upload POST.
* @param {String} photo_url - The photo url returned from the image upload POST
*/
classifyPhoto(photo_url) {
if (!this.ATTEMPTED_CLASSIFICATION) {
this.PHOTOS.push(photo_url);
const data = { "url": photo_url }
return new Promise((resove, reject) => {
this.makeRequest(this.REQUEST_TYPES.POST, this.AWS_ENDPOINT, data)
.then(data => {
const { label, confidence } = data;
this.PREDICTED_CLASS = label;
this.PREDICTED_CONFIDENCE = confidence;
this.ATTEMPTED_CLASSIFICATION = true;
resolve(data);
})
})
.catch(err => {
reject(err);
})
}
else {
return;
}
}Reflection
This time felt like a very successful use of our Professional Development time and resources. It was particularly rewarding to learn and advance our development craft, while making an impact with a local organization, and conservationists across the US and Canada.



